4 Input and Output
Introduction
All statistical work begins with data, and most data is stuck inside files and databases. Dealing with input is probably the first step of implementing any significant statistical project.
All statistical work ends with reporting numbers back to a client, even if you are the client. Formatting and producing output is probably the climax of your project.
Casual R users can solve their input problems by using basic readr package functions
such as read_csv to read CSV files and read_delim to read more
complicated, tabular data. They can use print, cat, and format to
produce simple reports.
Users with heavy-duty input/output (I/O) needs are strongly encouraged to read the R Data Import/Export guide, available on CRAN. This manual includes important information on reading data from sources such as spreadsheets, binary files, other statistical systems, and relational databases.
4.1 Entering Data from the Keyboard
Problem
You have a small amount of data, too small to justify the overhead of creating an input file. You just want to enter the data directly into your workspace.
Solution
For very small datasets, enter the data as literals using the c
constructor for vectors:
Discussion
When working on a simple problem, you may not want the hassle of creating
and then reading a data file outside of R. You may just want to enter the data into R.
The easiest way to do so is by using the c constructor for vectors, as shown
in the Solution.
You can use this approach for data frames, too, by entering each variable (column) as a vector:
See Also
For cutting and pasting data from another application into R, be sure to look at datapasta, a package that provides RStudio add-ins that make pasting data into your scripts easier.
4.2 Printing Fewer Digits (or More Digits)
Problem
Your output contains too many digits or too few digits. You want to print fewer or more.
Solution
For print, the digits parameter can control the number of printed
digits.
For cat, use the format function (which also has a digits
parameter) to alter the formatting of numbers.
Discussion
R normally formats floating-point output to have seven digits. This works well most of the time but can become annoying when you have lots of numbers to print in a small space. It gets downright misleading when there are only a few significant digits in your numbers and R still prints seven.
The print function lets you vary the number of printed digits using
the digits parameter:
The cat function does not give you direct control over formatting.
Instead, use the format function to format your numbers before calling
cat:
This is R, so both print and format will format entire vectors at
once:
print(pnorm(-3:3), digits = 2)
#> [1] 0.0013 0.0228 0.1587 0.5000 0.8413 0.9772 0.9987
format(pnorm(-3:3), digits = 2)
#> [1] "0.0013" "0.0228" "0.1587" "0.5000" "0.8413" "0.9772" "0.9987"Notice that both print and format format the vector elements consistently, finding
the number of significant digits necessary to format the smallest number and then
formatting all numbers to have the same width (though not necessarily
the same number of digits). This is extremely useful for formating an
entire table:
q <- seq(from = 0, to = 3, by = 0.5)
tbl <- data.frame(Quant = q,
Lower = pnorm(-q),
Upper = pnorm(q))
tbl # Unformatted print
#> Quant Lower Upper
#> 1 0.0 0.50000 0.500
#> 2 0.5 0.30854 0.691
#> 3 1.0 0.15866 0.841
#> 4 1.5 0.06681 0.933
#> 5 2.0 0.02275 0.977
#> 6 2.5 0.00621 0.994
#> 7 3.0 0.00135 0.999
print(tbl, digits = 2) # Formatted print: fewer digits
#> Quant Lower Upper
#> 1 0.0 0.5000 0.50
#> 2 0.5 0.3085 0.69
#> 3 1.0 0.1587 0.84
#> 4 1.5 0.0668 0.93
#> 5 2.0 0.0228 0.98
#> 6 2.5 0.0062 0.99
#> 7 3.0 0.0013 1.00As you can see, when an entire vector or column is formatted, each element in the vector or column is formatted the same way.
You can also alter the format of all output by using the options
function to change the default for digits:
But this is a poor choice in our experience, since it also alters the output from R’s built-in functions, and that alteration will likely be unpleasant.
See Also
Other functions for formatting numbers include sprintf and formatC;
see their help pages for details.
4.3 Redirecting Output to a File
Problem
You want to redirect the output from R into a file instead of your console.
Solution
You can redirect the output of the cat function by using its file
argument:
Use the sink function to redirect all output from both print and
cat. Call sink with a *filename* argument to begin redirecting console
output to that file. When you are done, use sink with no argument to
close the file and resume output to the console:
Discussion
The print and cat functions normally write their output to your
console. The cat function writes to a file if you supply a file
argument, which can be either a filename or a connection. The print
function cannot redirect its output, but the sink function can force all
output to a file. A common use for sink is to capture the output of an R
script:
sink("script_output.txt") # Redirect output to file
source("script.R") # Run the script, capturing its output
sink() # Resume writing output to consoleIf you are repeatedly cating items to one file, be sure to set
append=TRUE. Otherwise, each call to cat will simply overwrite the
file’s contents:
cat(data, file = "analysisReport.out")
cat(results, file = "analysisRepart.out", append = TRUE)
cat(conclusion, file = "analysisReport.out", append = TRUE)Hardcoding filenames like this is a tedious and error-prone process. Did you notice that the filename is misspelled in the second line? Instead of hardcoding the filename repeatedly, we suggest opening a connection to the file and writing your output to the connection:
con <- file("analysisReport.out", "w")
cat(data, file = con)
cat(results, file = con)
cat(conclusion, file = con)
close(con)(You don’t need append=TRUE when writing to a connection because append is the default with connections.) This technique is especially valuable inside R scripts because
it makes your code more reliable and more maintainable.
4.4 Listing Files
Problem
You want an R vector that is a listing of the files in your working directory.
Solution
The list.files function shows the contents of your working directory:
Discussion
This function is terribly handy to grab the names of all files in a subdirectory. You can use it to refresh your memory of your filenames or, more likely, as input into another process, like importing data files.
You can pass list.files a path and a pattern to shows files in a specific path and matching a specific regular expression pattern.
list.files(path = 'data/') # show files in a directory
#> [1] "ac.rdata" "adf.rdata"
#> [3] "anova.rdata" "anova2.rdata"
#> [5] "bad.rdata" "batches.rdata"
#> [7] "bnd_cmty.Rdata" "compositePerf-2010.csv"
#> [9] "conf.rdata" "daily.prod.rdata"
#> [11] "data1.csv" "data2.csv"
#> [13] "datafile_missing.tsv" "datafile.csv"
#> [15] "datafile.fwf" "datafile.qsv"
#> [17] "datafile.ssv" "datafile.tsv"
#> [19] "datafile1.ssv" "df_decay.rdata"
#> [21] "df_squared.rdata" "diffs.rdata"
#> [23] "example1_headless.csv" "example1.csv"
#> [25] "excel_table_data.xlsx" "get_USDA_NASS_data.R"
#> [27] "ibm.rdata" "iris_excel.xlsx"
#> [29] "lab_df.rdata" "movies.sas7bdat"
#> [31] "nacho_data.csv" "NearestPoint.R"
#> [33] "not_a_csv.txt" "opt.rdata"
#> [35] "outcome.rdata" "pca.rdata"
#> [37] "pred.rdata" "pred2.rdata"
#> [39] "sat.rdata" "singles.txt"
#> [41] "state_corn_yield.rds" "student_data.rdata"
#> [43] "suburbs.txt" "tab1.csv"
#> [45] "tls.rdata" "triples.txt"
#> [47] "ts_acf.rdata" "workers.rdata"
#> [49] "world_series.csv" "xy.rdata"
#> [51] "yield.Rdata" "z.RData"
list.files(path = 'data/', pattern = '\\.csv')
#> [1] "compositePerf-2010.csv" "data1.csv"
#> [3] "data2.csv" "datafile.csv"
#> [5] "example1_headless.csv" "example1.csv"
#> [7] "nacho_data.csv" "tab1.csv"
#> [9] "world_series.csv"To see all the files in your subdirectories, too, use:
A possible “gotcha” of list.files is that it ignores hidden
files—typically, any file whose name begins with a period. If you don’t
see the file you expected to see, try setting all.files=TRUE:
list.files(path = 'data/', all.files = TRUE)
#> [1] "." ".."
#> [3] ".hidden_file.txt" "ac.rdata"
#> [5] "adf.rdata" "anova.rdata"
#> [7] "anova2.rdata" "bad.rdata"
#> [9] "batches.rdata" "bnd_cmty.Rdata"
#> [11] "compositePerf-2010.csv" "conf.rdata"
#> [13] "daily.prod.rdata" "data1.csv"
#> [15] "data2.csv" "datafile_missing.tsv"
#> [17] "datafile.csv" "datafile.fwf"
#> [19] "datafile.qsv" "datafile.ssv"
#> [21] "datafile.tsv" "datafile1.ssv"
#> [23] "df_decay.rdata" "df_squared.rdata"
#> [25] "diffs.rdata" "example1_headless.csv"
#> [27] "example1.csv" "excel_table_data.xlsx"
#> [29] "get_USDA_NASS_data.R" "ibm.rdata"
#> [31] "iris_excel.xlsx" "lab_df.rdata"
#> [33] "movies.sas7bdat" "nacho_data.csv"
#> [35] "NearestPoint.R" "not_a_csv.txt"
#> [37] "opt.rdata" "outcome.rdata"
#> [39] "pca.rdata" "pred.rdata"
#> [41] "pred2.rdata" "sat.rdata"
#> [43] "singles.txt" "state_corn_yield.rds"
#> [45] "student_data.rdata" "suburbs.txt"
#> [47] "tab1.csv" "tls.rdata"
#> [49] "triples.txt" "ts_acf.rdata"
#> [51] "workers.rdata" "world_series.csv"
#> [53] "xy.rdata" "yield.Rdata"
#> [55] "z.RData"If you just want to see which files are in a directory and not use the filenames in a procedure, the easiest way is to open the Files pane in the lower-right corner of RStudio. But keep in mind that the RStudio Files pane hides files that start with a period, as you can see in Figure 4.1.
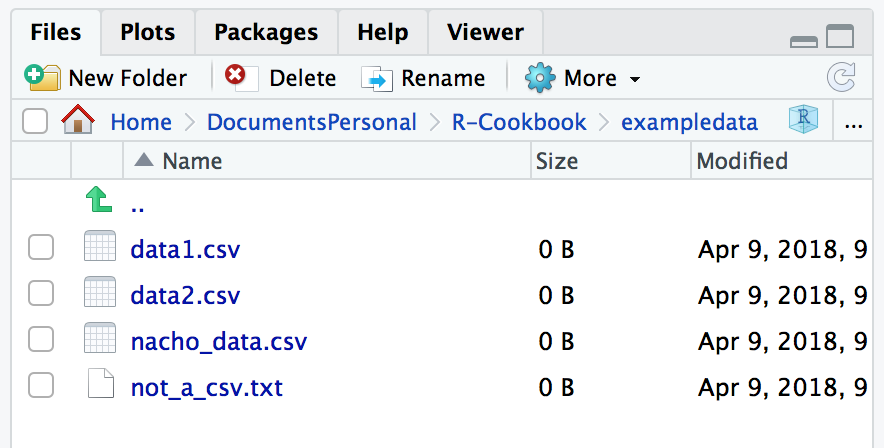
Figure 4.1: RStudio Files pane
See Also
R has other handy functions for working with files; see help(files).
4.5 Dealing with “Cannot Open File” in Windows
Problem
You are running R on Windows, and you are using filenames such as
C:\data\sample.txt. R says it cannot open the file, but you know the
file does exist.
Solution
The backslashes in the filepath are causing trouble. You can solve this problem in one of two ways:
Change the backslashes to forward slashes:
"C:/data/sample.txt".Double the backslashes:
"C:\\data\\sample.txt".
Discussion
When you open a file in R, you give the filename as a character string.
Problems arise when the name contains backslashes (\) because
backslashes have a special meaning inside strings. You’ll probably get
something like this:
samp <- read_csv("C:\Data\sample-data.csv")
#> Error: '\D' is an unrecognized escape in character string starting ""C:\D"R escapes every character that follows a backslash and then removes the
backslashes. That leaves a meaningless filepath, such as
C:Datasample-data.csv in this example.
The simple solution is to use forward slashes instead of backslashes. R leaves the forward slashes alone, and Windows treats them just like backslashes. Problem solved:
An alternative solution is to double the backslashes, since R replaces two consecutive backslashes with a single backslash:
4.6 Reading Fixed-Width Records
Problem
You are reading data from a file of fixed-width records: records whose data items occur at fixed boundaries.
Solution
Use the read_fwf from the readr package (which is part of the tidyverse). The main arguments are the filename and the description of the fields:
This form uses the fwf_cols parameter to pass column names and widths to the function. You can also pass column parameters in other ways as discussed next.
Discussion
For reading data into R, we highly recommend the readr package. There are Base R functions for reading in text files, but readr improves on these base functions
with faster performance, better defaults, and more flexibility.
Suppose we want to read an entire file of fixed-width records, such as fixed-width.txt, shown here:
Fisher R.A. 1890 1962
Pearson Karl 1857 1936
Cox Gertrude 1900 1978
Yates Frank 1902 1994
Smith Kirstine 1878 1939We need to know the column widths. In this case the columns are:
- Last name, 10 characters
- First name, 10 characters
- Year of birth, 5 characters
- Year of death, 5 characters
There are five different ways to define the columns using read_fwf. Pick the one that’s easiest to use (or remember) in your situation:
read_fwfcan try to guess your column widths if there is empty space between the columns, with thefwf_emptyoption:
file <- "./data/datafile.fwf"
t1 <- read_fwf(file, fwf_empty(file, col_names = c("last", "first", "birth", "death")))
#> Parsed with column specification:
#> cols(
#> last = col_character(),
#> first = col_character(),
#> birth = col_double(),
#> death = col_double()
#> )- You can define each column by a vector of widths followed by a vector of names with
fwf_widths:
t2 <- read_fwf(file, fwf_widths(c(10, 10, 5, 4),
c("last", "first", "birth", "death")))
#> Parsed with column specification:
#> cols(
#> last = col_character(),
#> first = col_character(),
#> birth = col_double(),
#> death = col_double()
#> )- The columns can be defined with
fwf_cols, which takes a series of column names followed by the column widths:
t3 <-
read_fwf("./data/datafile.fwf",
fwf_cols(
last = 10,
first = 10,
birth = 5,
death = 5
))
#> Parsed with column specification:
#> cols(
#> last = col_character(),
#> first = col_character(),
#> birth = col_double(),
#> death = col_double()
#> )- Each column can be defined by a beginning position and ending position with
fwf_cols:
t4 <- read_fwf(file, fwf_cols(
last = c(1, 10),
first = c(11, 20),
birth = c(21, 25),
death = c(26, 30)
))
#> Parsed with column specification:
#> cols(
#> last = col_character(),
#> first = col_character(),
#> birth = col_double(),
#> death = col_double()
#> )- You can also define the columns with a vector of starting positions, a vector of ending positions, and a vector of column names, with
fwf_positions:
t5 <- read_fwf(file, fwf_positions(
c(1, 11, 21, 26),
c(10, 20, 25, 30),
c("first", "last", "birth", "death")
))
#> Parsed with column specification:
#> cols(
#> first = col_character(),
#> last = col_character(),
#> birth = col_double(),
#> death = col_double()
#> )The read_fwf returns a tibble, which is a tidyverse flavor of data frame. As is common with tidyverse packages, read_fwf has a good selection of default assumptions that make it less tricky to use than some Base R functions for importing data. For example, read_fwf will, by default, import character fields as characters, not factors, which prevents much pain and consternation for users.
See Also
See Recipe 4.7, “Reading Tabular Data Files”, for more discussion of reading text files.
4.7 Reading Tabular Data Files
Problem
You want to read a text file that contains a table of whitespace-delimited data.
Solution
Use the read_table2 function from the readr package, which returns a tibble:
library(tidyverse)
tab1 <- read_table2("./data/datafile.tsv")
#> Parsed with column specification:
#> cols(
#> last = col_character(),
#> first = col_character(),
#> birth = col_double(),
#> death = col_double()
#> )
tab1
#> # A tibble: 5 x 4
#> last first birth death
#> <chr> <chr> <dbl> <dbl>
#> 1 Fisher R.A. 1890 1962
#> 2 Pearson Karl 1857 1936
#> 3 Cox Gertrude 1900 1978
#> 4 Yates Frank 1902 1994
#> 5 Smith Kirstine 1878 1939Discussion
Tabular data files are quite common. They are text files with a simple format:
Each line contains one record.
Within each record, fields (items) are separated by a whitespace delimiter, such as a space or tab.
Each record contains the same number of fields.
This format is more free-form than the fixed-width format because fields needn’t be aligned by position. Here is the data file from Recipe 4.6, “Reading Fixed-Width Records” in tabular format, using a tab character between fields:
last first birth death
Fisher R.A. 1890 1962
Pearson Karl 1857 1936
Cox Gertrude 1900 1978
Yates Frank 1902 1994
Smith Kirstine 1878 1939
The read_table2 function is designed to make some good guesses about your data. It assumes your data has column names in the first row, it guesses your delimiter, and it imputes your column types based on the first 1,000 records in your dataset. Next is an example with space-delimited data.
The source file looks like this:
last first birth death
Fisher R.A. 1890 1962
Pearson Karl 1857 1936
Cox Gertrude 1900 1978
Yates Frank 1902 1994
Smith Kirstine 1878 1939And read_table2 makes some rational guesses:
t <- read_table2("./data/datafile1.ssv")
#> Parsed with column specification:
#> cols(
#> last = col_character(),
#> first = col_character(),
#> birth = col_double(),
#> death = col_double()
#> )
print(t)
#> # A tibble: 5 x 4
#> last first birth death
#> <chr> <chr> <dbl> <dbl>
#> 1 Fisher R.A. 1890 1962
#> 2 Pearson Karl 1857 1936
#> 3 Cox Gertrude 1900 1978
#> 4 Yates Frank 1902 1994
#> 5 Smith Kirstine 1878 1939read_table2 often guesses correctly. But as with other readr import functions, you can overwrite the defaults with explicit parameters.
t <-
read_table2(
"./data/datafile1.ssv",
col_types = c(
col_character(),
col_character(),
col_integer(),
col_integer()
)
)If any field contains the string "NA", then read_table2 assumes that
the value is missing and converts it to NA. Your data file might employ
a different string to signal missing values, in which case use the
na parameter. The SAS convention, for example, is that missing
values are signaled by a single period (.).
We can read such text files using the na="." option.
If we have a file named datafile_missing.tsv that has a missing value indicated with a . in the last row:
last first birth death
Fisher R.A. 1890 1962
Pearson Karl 1857 1936
Cox Gertrude 1900 1978
Yates Frank 1902 1994
Smith Kirstine 1878 1939
Cox David 1924 .we can import it like so:
t <- read_table2("./data/datafile_missing.tsv", na = ".")
#> Parsed with column specification:
#> cols(
#> last = col_character(),
#> first = col_character(),
#> birth = col_double(),
#> death = col_double()
#> )
t
#> # A tibble: 6 x 4
#> last first birth death
#> <chr> <chr> <dbl> <dbl>
#> 1 Fisher R.A. 1890 1962
#> 2 Pearson Karl 1857 1936
#> 3 Cox Gertrude 1900 1978
#> 4 Yates Frank 1902 1994
#> 5 Smith Kirstine 1878 1939
#> 6 Cox David 1924 NAWe’re huge fans of self-describing data: data files that describe their
own contents.
(A computer scientist would say the file contains its own metadata.)
The read_table2 function makes the default assumption that the first line of your file contains a header line with column names.
If your file does not have column names, you can turn this off with the parameter col_names = FALSE.
An additional type of metadata supported by read_table2 is comment lines. Using the comment parameter you can tell read_table2 which character distinguishes comment lines. The following file has a comment line at the top that starts with #.
# The following is a list of statisticians
last first birth death
Fisher R.A. 1890 1962
Pearson Karl 1857 1936
Cox Gertrude 1900 1978
Yates Frank 1902 1994
Smith Kirstine 1878 1939so we can import this file as follows:
t <- read_table2("./data/datafile.ssv", comment = '#')
#> Parsed with column specification:
#> cols(
#> last = col_character(),
#> first = col_character(),
#> birth = col_double(),
#> death = col_double()
#> )
t
#> # A tibble: 5 x 4
#> last first birth death
#> <chr> <chr> <dbl> <dbl>
#> 1 Fisher R.A. 1890 1962
#> 2 Pearson Karl 1857 1936
#> 3 Cox Gertrude 1900 1978
#> 4 Yates Frank 1902 1994
#> 5 Smith Kirstine 1878 1939read_table2 has many parameters for controlling how it reads and
interprets the input file. See the help page (?read_table2) or the readr vignette (vignette("readr")) for more details. If you’re curious about the difference between read_table and read_table2, it’s in the help file… but the short answer is that read_table is slightly less forgiving in file structure and line length.
See Also
If your data items are separated by commas, see Recipe 4.8, “Reading from CSV Files”, for reading a CSV file.
4.8 Reading from CSV Files
Problem
You want to read data from a comma-separated values (CSV) file.
Solution
The read_csv function from the readr package is a fast (and, according to the documentation, fun) way to read CSV files. If your CSV file has a
header line, use this:
If your CSV file does not contain a header line, set the col_names option
to FALSE:
Discussion
The CSV file format is popular because many programs can import and export data in that format. This includes R, Excel, other spreadsheet programs, many database managers, and most statistical packages. It is a flat file of tabular data, where each line in the file is a row of data, and each row contains data items separated by commas. Here is a very simple CSV file with three rows and three columns. The first line is a header line that contains the column names, also separated by commas:
label,lbound,ubound
low,0,0.674
mid,0.674,1.64
high,1.64,2.33The read_csv file reads the data and creates a tibble. The function assumes that your file has a header line unless told otherwise:
tbl <- read_csv("./data/example1.csv")
#> Parsed with column specification:
#> cols(
#> label = col_character(),
#> lbound = col_double(),
#> ubound = col_double()
#> )
tbl
#> # A tibble: 3 x 3
#> label lbound ubound
#> <chr> <dbl> <dbl>
#> 1 low 0 0.674
#> 2 mid 0.674 1.64
#> 3 high 1.64 2.33Observe that read_csv took the column names from the header line for
the tibble. If the file did not contain a header, then we would
specify col_names=FALSE and R would synthesize column names for us (X1,
X2, and X3 in this case):
tbl <- read_csv("./data/example1.csv", col_names = FALSE)
#> Parsed with column specification:
#> cols(
#> X1 = col_character(),
#> X2 = col_character(),
#> X3 = col_character()
#> )
tbl
#> # A tibble: 4 x 3
#> X1 X2 X3
#> <chr> <chr> <chr>
#> 1 label lbound ubound
#> 2 low 0 0.674
#> 3 mid 0.674 1.64
#> 4 high 1.64 2.33Sometimes it’s convenient to put metadata in files. If this metadata starts with a common character, such as a pound sign (#), we can use the comment=FALSE parameter to ignore metadata lines.
The read_csv function has many useful bells and whistles. A few of these options and their default values include:
na = c("", "NA")Indicates what values represent missing or NA values
comment = ""Indicates which lines to ignore as comments or metadata
trim_ws = TRUEIndicates whether to drop whitespace at the beginning and/or end of fields
skip = 0Indicates the number of rows to skip at the beginning of the file
guess_max = min(1000, n_max)Indicates the number of rows to consider when imputing column types
See the R help page, help(read_csv), for more details on all the available options.
If you have a data file that uses semicolons (;) for separators and commas for the decimal mark, as is common outside of North America, you should use the function read_csv2, which is built for that very situation.
See Also
See Recipe 4.9, “Writing to CSV Files”. See also the vignette for the readr: vignette(readr).
4.9 Writing to CSV Files
Problem
You want to save a matrix or data frame in a file using the comma-separated values format.
Solution
The write_csv function from the tidyverse readr package can write a CSV file:
Discussion
The write_csv function writes tabular data to an ASCII file in CSV format. Each row of data creates one line in the file, with data items separated by commas (,). We can start with the data frame tab1 we created previously in Recipe 4.7, “Reading Tabular Data Files”.
This example creates a file called tab1.csv in the data directory, which is a subdirectory of the current working directory. The file looks like this:
last,first,birth,death
Fisher,R.A.,1890,1962
Pearson,Karl,1857,1936
Cox,Gertrude,1900,1978
Yates,Frank,1902,1994
Smith,Kirstine,1878,1939write_csv has a number of parameters with typically very good defaults. Should you want to adjust the output, here are a few parameters you can change, along with their defaults:
col_names = TRUEIndicates whether or not the first row contains column names.
col_types = NULLwrite_csvwill look at the first 1,000 rows (changeable withguess_max) and make an informed guess as to what data types to use for the columns. If you’d rather explicitly state the column types, you can do so by passing a vector of column types to the parametercol_types.na = c("", "NA")Indicates what values represent missing or NA values.
comment = ""Indicates which lines to ignore as comments or metadata.
trim_ws = TRUEIndicates whether to drop whitespace at the beginning and/or end of fields.
skip = 0Indicates the number of rows to skip at the beginning of the file.
guess_max = min(1000, n_max)Indicates the number of rows to consider when guessing column types.
See Also
See Recipe 3.1, “Getting and Setting the Working Directory”, for more about the current working directory
and Recipe 4.18, “Saving and Transporting Objects”, for other ways to save data to files. For more info on reading and writing text files, see the readr vignette: vignette(readr).
4.10 Reading Tabular or CSV Data from the Web
Problem
You want to read data directly from the web into your R workspace.
Solution
Use the read_csv or read_table2 functions from the readr package,
using a URL instead of a filename. The functions will read directly from the remote server:
library(tidyverse)
berkley <- read_csv('http://bit.ly/barkley18', comment = '#')
#> Parsed with column specification:
#> cols(
#> Name = col_character(),
#> Location = col_character(),
#> Time = col_time(format = "")
#> )You can also open a connection using the URL and then read from the connection, which may be preferable for complicated files.
Discussion
The web is a gold mine of data. You could download the data into a file
and then read the file into R, but it’s more convenient to read directly
from the web. Give the URL to read_csv, read_table2, or another read function in readr
(depending upon the format of the data), and the data will be downloaded
and parsed for you. No fuss, no muss.
Aside from using a URL, this recipe is just like reading from a CSV file (see Recipe 4.8, “Reading from CSV Files”) or a complex file (Recipe 4.15, “Reading Files with a Complex Structure”), so all the comments in those recipes apply here, too.
Remember that URLs work for FTP servers, not just HTTP servers. This means that R can also read data from FTP sites using URLs:
See Also
See Recipes 4.8, “Reading from CSV Files”, and 4.15, “Reading Files with a Complex Structure”.
4.11 Reading Data from Excel
Problem
You want to read data in from an Excel file.
Solution
The openxlsx package makes reading Excel files easy.
Discussion
The package openxlsx is a good choice for both reading and writing Excel files with R. If we’re reading in an entire sheet, passing a filename and a sheet name to the read.xlsx function is a simple option. We need only pass in a filename and, if desired, the name of the sheet we want imported:
library(openxlsx)
df1 <- read.xlsx(xlsxFile = "data/iris_excel.xlsx",
sheet = 'iris_data')
head(df1, 3)
#> Sepal.Length Sepal.Width Petal.Length Petal.Width Species
#> 1 5.1 3.5 1.4 0.2 setosa
#> 2 4.9 3.0 1.4 0.2 setosa
#> 3 4.7 3.2 1.3 0.2 setosaBut openxlsx supports more complex workflows.
A common pattern is to read a named table out of an Excel file and into an R data frame. This is trickier because the sheet we’re reading from may have values outside of the named table and we want to only read in the named table range. We can use the functions in openxlsx to get the location of a table, then read that range of cells into a data frame.
First we load the entire workbook into R:
Then we can use the getTables function to get the names and ranges of all the Excel tables in the input_data sheet and select out the one table we want. In this example the Excel table we are after is named example_data:
tables <- getTables(wb, 'input_data')
table_range_str <- names(tables[tables == 'example_table'])
table_range_refs <- strsplit(table_range_str, ':')[[1]]
# use a regex to extract out the row numbers
table_range_row_num <- gsub("[^0-9.]", "", table_range_refs)
# extract out the column numbers
table_range_col_num <- convertFromExcelRef(table_range_refs)Now the vector table_range_col_num contains the column numbers of our named table, while table_range_row_num contains the row numbers.
We can then use the read.xlsx function to pull in only the rows and columns we want.
df <- read.xlsx(
xlsxFile = "data/excel_table_data.xlsx",
sheet = 'input_data',
cols = table_range_col_num[1]:table_range_col_num[2],
rows = table_range_row_num[1]:table_range_row_num[2]
)While this may seem complicated, this design pattern can save a lot of hassle when sharing data with analysts who are using highly structured Excel files that include named tables.
See Also
Vignette for openxlsx by installing openxlsx and running: vignette('Introduction', package='openxlsx')
The readxl package is part of the tidyverse and provides fast, simple reading of Excel files. However, readxl does not currently support named Excel tables.
The writexl package is a fast and lightweight (no dependencies) package for writing Excel files.
Recipe 4.12, “Writing a Data Frame to Excel”
4.12 Writing a Data Frame to Excel
Problem
You want to write an R data frame to an Excel file.
Solution
The openxlsx package makes writing to Excel files relatively easy. While there are lots of options in openxlsx, a typical pattern is to specify an Excel filename and a sheet name:
Discussion
The openxlsx package has a huge number of options for controlling many aspects of the Excel object model. We can use it to set cell colors, define named ranges, and set cell outlines, for example. But it has a few helper functions like write.xlsx that make simple tasks super easy.
When businesses work with Excel, it’s a good practice to keep all input data in an Excel file in a named Excel table, which makes accessing the data easier and less error prone. However, if you use openxlsx to overwrite an Excel table in one of the sheets, you run the risk that the new data may contain fewer rows than the Excel table it replaces. That could cause errors, as you would end up with old data and new data in contiguous rows. The solution is to first delete an existing Excel table, then add new data back into the same location and assign the new data to a named Excel table. To do this we need to use the more advanced Excel manipulation features of openxlsx.
First we use loadWorkbook to read the Excel workbook into R in its entirety:
Before we delete the table, we want to extract the table’s starting row and column.
tables <- getTables(wb, 'input_data')
table_range_str <- names(tables[tables == 'example_table'])
table_range_refs <- strsplit(table_range_str, ':')[[1]]
# use a regex to extract out the starting row number
table_row_num <- gsub("[^0-9.]", "", table_range_refs)[[1]]
# extract out the starting column number
table_col_num <- convertFromExcelRef(table_range_refs)[[1]]Then we can use the removeTable function to remove the existing named Excel table:
Then we can use writeDataTable to write the iris data frame (which comes with R) to write data back into our workbook object in R.
writeDataTable(
wb = wb,
sheet = 'input_data',
x = iris,
startCol = table_col_num,
startRow = table_row_num,
tableStyle = "TableStyleLight9",
tableName = 'example_table'
)At this point we could save the workbook and our table would be updated. However, it’s a good idea to save some metadata in the workbook to let others know exactly when the data was refreshed. We can do this with the writeData function, then save the workbook to a file and overwrite the original file. In this example, we’ll put the metadata text in cell B:5, then save the workbook back to a file, overwriting the original.
writeData(
wb = wb,
sheet = 'input_data',
x = paste('example_table data refreshed on:', Sys.time()),
startCol = 2,
startRow = 5
)
## then save the workbook
saveWorkbook(wb = wb,
file = "data/excel_table_data.xlsx",
overwrite = TRUE)
#> Note: zip::zip() is deprecated, please use zip::zipr() insteadThe resulting Excel sheet looks as shown in Figure 4.2.
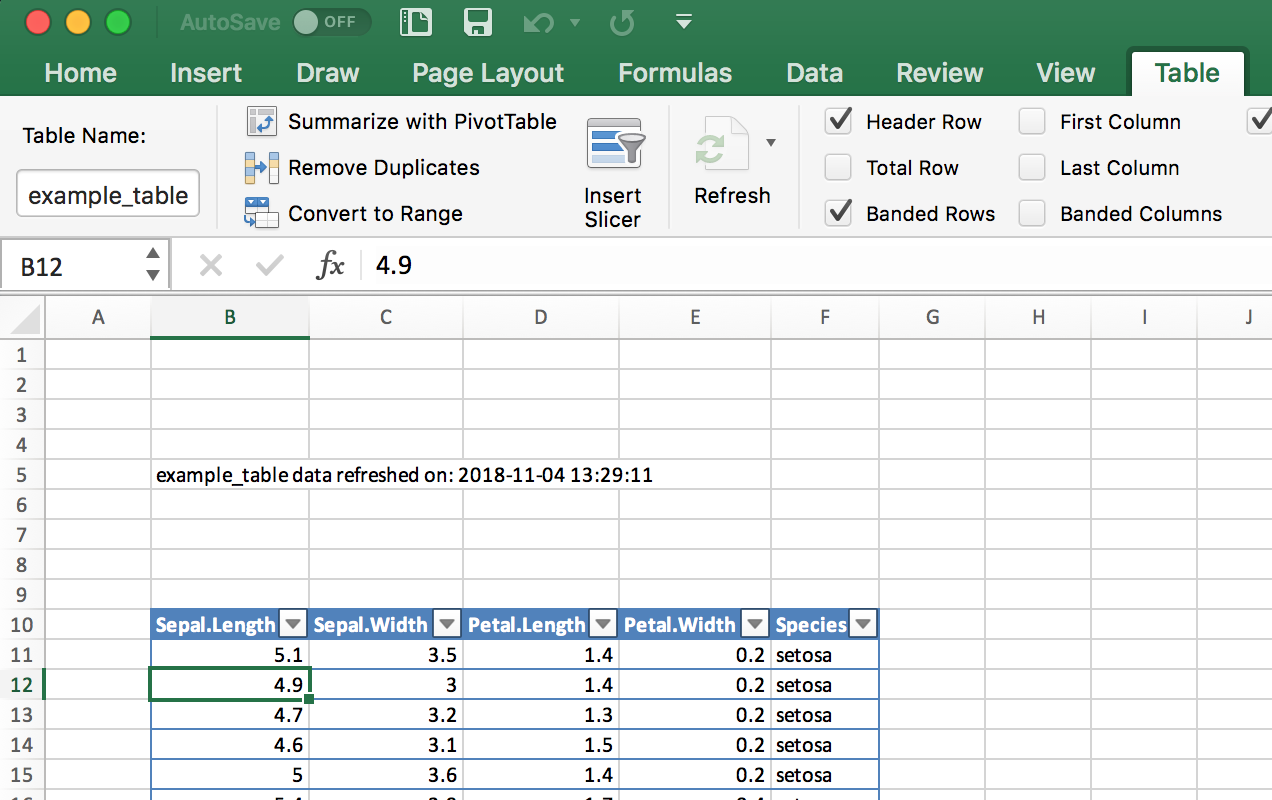
Figure 4.2: Excel table and metadata text
See Also
Vignette for openxlsx by installing openxlsx and running: vignette('Introduction', package='openxlsx')
The readxl package is part of the tidyverse and provides fast, simple reading of Excel files.
The writexl package is a fast and lightweight (no dependencies) package for writing Excel files.
Recipe 4.11, “Reading Data from Excel”
4.13 Reading Data from a SAS file
Problem
You want to read a SAS dataset into an R data frame.
Solution
The sas7bdat package supports reading SAS sas7bdat files into R.
Discussion
SAS V7 and beyond all support the sas7bdat file format. The read_sas function in haven supports reading the sas7bdat file format including variable labels. If your SAS file has variable labels, when they are inported into R they will be stored in the label attributes of the data frame. These labels will not be printed by default. You can see the labels by opening the data frame in RStudio, or by calling the attributes Base R function on each column:
sapply(sas_movie_data, attributes)
#> $Movie
#> $Movie$label
#> [1] "Movie"
#>
#>
#> $Type
#> $Type$label
#> [1] "Type"
#>
#>
#> $Rating
#> $Rating$label
#> [1] "Rating"
#>
#>
#> $Year
#> $Year$label
#> [1] "Year"
#>
#>
#> $Domestic__
#> $Domestic__$label
#> [1] "Domestic $"
#>
#> $Domestic__$format.sas
#> [1] "F"
#>
#>
#> $Worldwide__
#> $Worldwide__$label
#> [1] "Worldwide $"
#>
#> $Worldwide__$format.sas
#> [1] "F"
#>
#>
#> $Director
#> $Director$label
#> [1] "Director"See Also
The sas7bdat package is much slower on large files than haven, but it has more elaborate support for file attributes. If the SAS metadata is important to you, then you should investigate sas7bdat::read.sas7bdat.
4.14 Reading Data from HTML Tables
Problem
You want to read data from an HTML table on the web.
Solution
Use the read_html and html_table functions in the rvest package. To read all
tables on the page, do the following:
library(rvest)
library(tidyverse)
all_tables <-
read_html("URL") %>%
html_table(fill = TRUE, header = TRUE)Note that rvest is installed when you run install.packages('tidyverse'), although it is not a core tidyverse package. So you must explicitly load the package.
Discussion
Web pages can contain several HTML tables. Calling read_html(*url*) and then piping that to html_table
reads all tables on the page and returns them in a list. This can be
useful for exploring a page, but it’s annoying if you want just one
specific table. In that case, use extract2(*n*) to select the nth table.
Here we can extract all tables from a Wikipedia article:
library(rvest)
all_tables <-
read_html("https://en.wikipedia.org/wiki/Aviation_accidents_and_incidents") %>%
html_table(fill = TRUE, header = TRUE)read_html puts all tables from the HTML document into the output list. To pull a single table from that list, you can use the function extract2 from the magrittr package:
out_table <-
all_tables %>%
magrittr::extract2(2)
head(out_table)
#> Year Deaths[55] # of incidents[56]
#> 1 2018 1,040 113[57]
#> 2 2017 399 101
#> 3 2016 629 102
#> 4 2015 898 123
#> 5 2014 1,328 122
#> 6 2013 459 138Two common parameters for the html_table function are fill=TRUE, which fills in missing values with NA, and header=TRUE, which indicates that the first row contains the header names.
The following example loads all tables from the Wikipedia page entitled “World population”:
url <- 'http://en.wikipedia.org/wiki/World_population'
tbls <- read_html(url) %>%
html_table(fill = TRUE, header = TRUE)As it turns out, that page contains 23 tables (or things that html_table thinks might be tables):
In this example we care only about the sixth table (which lists the
largest populations by country), so we can either access that element using brackets, tbls[[6]], or we can pipe it into the extract2 function from the magrittr package:
library(magrittr)
tbl <- tbls %>%
extract2(6)
head(tbl, 2)
#> Rank Country / Territory Population Date
#> 1 1 China[note 4] 1,399,270,000 September 25, 2019
#> 2 2 India 1,352,730,000 September 25, 2019
#> % of worldpopulation Source
#> 1 18.1% [85]
#> 2 17.5% [86]The extract2 function is a “pipe friendly” version of the R [[i]] syntax. So it pulls out a single list element from a list. The extract function is analogous to [i], which returns element i from the original list into a list of length 1.
In that table, columns 2 and 3 contain the country name and population, respectively:
tbl[, c(2, 3)]
#> Country / Territory Population
#> 1 China[note 4] 1,399,270,000
#> 2 India 1,352,730,000
#> 3 United States 329,957,000
#> 4 Indonesia 265,015,300
#> 5 Pakistan 212,742,631
#> 6 Brazil 210,485,000
#> 7 Nigeria 188,500,000
#> 8 Bangladesh 167,303,000
#> 9 Russia[note 5] 146,877,088
#> 10 Mexico 126,577,691Right away, we can see problems with the data: China and Russia have [note 4] and [note 5] appended to their names. On the
Wikipedia website, that was a footnote reference, but now it’s just a
bit of unwanted text. Adding insult to injury, the population numbers
have embedded commas, so you cannot easily convert them to raw numbers.
All these problems can be solved by some string processing, but each
problem adds at least one more step to the process.
This illustrates the main obstacle to reading HTML tables. HTML was designed for presenting information to people, not to computers. When you “scrape” information off an HTML page, you get stuff that’s useful to people but annoying to computers. If you ever have a choice, choose instead a computer-oriented data representation such as XML, JSON, or CSV.
The read_html(*url*) and html_table functions are part of the rvest package, which (by necessity) is large and complex. Any time you pull data from a site designed for human readers, not machines, expect that you will have to do post-processing to clean up the bits and pieces the machine leaves messy.
See Also
See Recipe 3.10, “Installing Packages from CRAN”, for downloading and installing packages such as
the rvest package.
4.15 Reading Files with a Complex Structure
Problem
You are reading data from a file that has a complex or irregular structure.
Solution
Use the
readLinesfunction to read individual lines; then process them as strings to extract data items.Alternatively, use the
scanfunction to read individual tokens and use the argumentwhatto describe the stream of tokens in your file. The function can convert tokens into data and then assemble the data into records.
Discussion
Life would be simple and beautiful if all our data files were organized
into neat tables with cleanly delimited data. We could read those files
using one of the functions in the readr package and get on with living.
Unfortunately, we don’t live in a land of rainbows and unicorn kisses.
You will eventually encounter a funky file format, and your job is to read the file contents into R.
The read.table and read.csv functions are file-oriented and probably won’t help.
However, the readLines and scan functions are useful here because
they let you process the individual lines and even tokens of the file.
The readLines function is pretty simple. It reads lines from a file
and returns them as a list of character strings:
You can limit the number of lines by using the n parameter, which
gives the maximum number of lines to be read:
The scan function is much richer. It reads one token at a time and
handles it according to your instructions. The first argument is either
a filename or a connection. The second
argument is called what, and it describes the tokens that scan
should expect in the input file. The description is cryptic but quite
clever:
what=numeric(0)Interprets the next token as a number.
what=integer(0)Interprets the next token as an integer.
what=complex(0)Interprets the next token as a complex number.
what=character(0)Interprets the next token as a character string.
what=logical(0)Interprets the next token as a logical value.
The scan function will apply the given pattern repeatedly until all
data is read.
Suppose your file is simply a sequence of numbers, like this:
2355.09 2246.73 1738.74 1841.01 2027.85Use what=numeric(0) to say, “My file is a sequence of tokens, each of
which is a number”:
singles <- scan("./data/singles.txt", what = numeric(0))
singles
#> [1] 2355.09 2246.73 1738.74 1841.01 2027.85A key feature of scan is that the what can be a list containing
several token types. The scan function will assume your file is a
repeating sequence of those types. Suppose your file contains triplets
of data, like this:
15-Oct-87 2439.78 2345.63 16-Oct-87 2396.21 2207.73
19-Oct-87 2164.16 1677.55 20-Oct-87 2067.47 1616.21
21-Oct-87 2081.07 1951.76Use a list to tell scan that it should expect a repeating, three-token
sequence:
triples <-
scan("./data/triples.txt",
what = list(character(0), numeric(0), numeric(0)))
triples
#> [[1]]
#> [1] "15-Oct-87" "16-Oct-87" "19-Oct-87" "20-Oct-87" "21-Oct-87"
#>
#> [[2]]
#> [1] 2439.78 2396.21 2164.16 2067.47 2081.07
#>
#> [[3]]
#> [1] 2345.63 2207.73 1677.55 1616.21 1951.76Give names to the list elements, and scan will assign those names to
the data:
triples <- scan("./data/triples.txt",
what = list(
date = character(0),
high = numeric(0),
low = numeric(0)
))
triples
#> $date
#> [1] "15-Oct-87" "16-Oct-87" "19-Oct-87" "20-Oct-87" "21-Oct-87"
#>
#> $high
#> [1] 2439.78 2396.21 2164.16 2067.47 2081.07
#>
#> $low
#> [1] 2345.63 2207.73 1677.55 1616.21 1951.76This can easily be turned into a data frame with the data.frame command:
df_triples <- data.frame(triples)
df_triples
#> date high low
#> 1 15-Oct-87 2439.78 2345.63
#> 2 16-Oct-87 2396.21 2207.73
#> 3 19-Oct-87 2164.16 1677.55
#> 4 20-Oct-87 2067.47 1616.21
#> 5 21-Oct-87 2081.07 1951.76The scan function has many bells and whistles, but the following are
especially useful:
n=numberStop after reading this many tokens. (Default: stop at end of file.)
nlines=numberStop after reading this many input lines. (Default: stop at end of file.)
skip=numberNumber of input lines to skip before reading data.
na.strings=listA list of strings to be interpreted as NA.
An Example
Let’s use this recipe to read a dataset from StatLib, the repository of
statistical data and software maintained by Carnegie Mellon University.
Jeff Witmer contributed a dataset called wseries that shows the
pattern of wins and losses for every World Series since 1903. The
dataset is stored in an ASCII file with 35 lines of comments followed by
23 lines of data. The data itself looks like this:
1903 LWLlwwwW 1927 wwWW 1950 wwWW 1973 WLwllWW
1905 wLwWW 1928 WWww 1951 LWlwwW 1974 wlWWW
1906 wLwLwW 1929 wwLWW 1952 lwLWLww 1975 lwWLWlw
1907 WWww 1930 WWllwW 1953 WWllwW 1976 WWww
1908 wWLww 1931 LWwlwLW 1954 WWww 1977 WLwwlW
.
. (etc.)
. The data is encoded as follows: L = loss at home, l = loss on the road, W = win at home, w = win on the road. The data appears in column order, not row order, which complicates our lives a bit.
Here is the R code for reading the raw data:
# Read the wseries dataset:
# - Skip the first 35 lines
# - Then read 23 lines of data
# - The data occurs in pairs: a year and a pattern (char string)
#
world.series <- scan(
"http://lib.stat.cmu.edu/datasets/wseries",
skip = 35,
nlines = 23,
what = list(year = integer(0),
pattern = character(0)),
)The scan function returns a list, so we get a list with two elements:
year and pattern. The function reads from left to right, but the
dataset is organized by columns and so the years appear in a strange
order:
world.series$year
#> [1] 1903 1927 1950 1973 1905 1928 1951 1974 1906 1929 1952 1975 1907 1930
#> [15] 1953 1976 1908 1931 1954 1977 1909 1932 1955 1978 1910 1933 1956 1979
#> [29] 1911 1934 1957 1980 1912 1935 1958 1981 1913 1936 1959 1982 1914 1937
#> [43] 1960 1983 1915 1938 1961 1984 1916 1939 1962 1985 1917 1940 1963 1986
#> [57] 1918 1941 1964 1987 1919 1942 1965 1988 1920 1943 1966 1989 1921 1944
#> [71] 1967 1990 1922 1945 1968 1991 1923 1946 1969 1992 1924 1947 1970 1993
#> [85] 1925 1948 1971 1926 1949 1972We can fix that by sorting the list elements according to year:
perm <- order(world.series$year)
world.series <- list(year = world.series$year[perm],
pattern = world.series$pattern[perm])Now the data appears in chronological order:
world.series$year
#> [1] 1903 1905 1906 1907 1908 1909 1910 1911 1912 1913 1914 1915 1916 1917
#> [15] 1918 1919 1920 1921 1922 1923 1924 1925 1926 1927 1928 1929 1930 1931
#> [29] 1932 1933 1934 1935 1936 1937 1938 1939 1940 1941 1942 1943 1944 1945
#> [43] 1946 1947 1948 1949 1950 1951 1952 1953 1954 1955 1956 1957 1958 1959
#> [57] 1960 1961 1962 1963 1964 1965 1966 1967 1968 1969 1970 1971 1972 1973
#> [71] 1974 1975 1976 1977 1978 1979 1980 1981 1982 1983 1984 1985 1986 1987
#> [85] 1988 1989 1990 1991 1992 1993
world.series$pattern
#> [1] "LWLlwwwW" "wLwWW" "wLwLwW" "WWww" "wWLww" "WLwlWlw"
#> [7] "WWwlw" "lWwWlW" "wLwWlLW" "wLwWw" "wwWW" "lwWWw"
#> [13] "WWlwW" "WWllWw" "wlwWLW" "WWlwwLLw" "wllWWWW" "LlWwLwWw"
#> [19] "WWwW" "LwLwWw" "LWlwlWW" "LWllwWW" "lwWLLww" "wwWW"
#> [25] "WWww" "wwLWW" "WWllwW" "LWwlwLW" "WWww" "WWlww"
#> [31] "wlWLLww" "LWwwlW" "lwWWLw" "WWwlw" "wwWW" "WWww"
#> [37] "LWlwlWW" "WLwww" "LWwww" "WLWww" "LWlwwW" "LWLwwlw"
#> [43] "LWlwlww" "WWllwLW" "lwWWLw" "WLwww" "wwWW" "LWlwwW"
#> [49] "lwLWLww" "WWllwW" "WWww" "llWWWlw" "llWWWlw" "lwLWWlw"
#> [55] "llWLWww" "lwWWLw" "WLlwwLW" "WLwww" "wlWLWlw" "wwWW"
#> [61] "WLlwwLW" "llWWWlw" "wwWW" "wlWWLlw" "lwLLWww" "lwWWW"
#> [67] "wwWLW" "llWWWlw" "wwLWLlw" "WLwllWW" "wlWWW" "lwWLWlw"
#> [73] "WWww" "WLwwlW" "llWWWw" "lwLLWww" "WWllwW" "llWWWw"
#> [79] "LWwllWW" "LWwww" "wlWWW" "LLwlwWW" "LLwwlWW" "WWlllWW"
#> [85] "WWlww" "WWww" "WWww" "WWlllWW" "lwWWLw" "WLwwlW"4.16 Reading from MySQL Databases
Problem
You want access to data stored in a MySQL database.
Solution
Install the
RMySQLpackage on your computer and add a user and password.Open a database connection using the
DBI::dbConnectfunction.Use
dbGetQueryto initiate aSELECTand return the result sets.Use
dbDisconnectto terminate the database connection when you are done.
Discussion
This recipe requires that the RMySQL package be installed on your
computer. That package requires, in turn, the MySQL client software. If
the MySQL client software is not already installed and configured,
consult the MySQL documentation or your system administrator.
Use the dbConnect function to establish a connection to the MySQL
database. It returns a connection object that is used in subsequent
calls to RMySQL functions:
library(RMySQL)
con <- dbConnect(
drv = RMySQL::MySQL(),
dbname = "your_db_name",
host = "your.host.com",
username = "userid",
password = "pwd"
)The username, password, and host parameters are the same parameters used
for accessing MySQL through the mysql client program. The example
given here shows them hardcoded into the dbConnect call. Actually,
that is an ill-advised practice. It puts your password in a plain-text document,
creating a security problem. It also creates a major headache whenever
your password or host changes, requiring you to hunt down the hardcoded
values. We strongly recommend using the security mechanism of MySQL
instead. Version 8 of MySQL introduces even more advanced security options, but currently these have not been built into the R MySQL client. So we recommend you use MySQL native passwords by setting default-authentication-plugin=mysql_native_password in your MySQL configuration file,
which is $HOME/.my.cnf on Unix and C:\my.cnf on Windows. We use loose-local-infile=1 to ensure that we have permissions to write to the database. Make sure the file is unreadable by anyone except you. The file is delimited into
sections with markers such as [mysqld] and [client]. Put connection parameters into the
[client] section, so that your config file will contain something like
this:
[mysqld]
default-authentication-plugin=mysql_native_password
loose-local-infile=1
[client]
loose-local-infile=1
user="jdl"
password="password"
host=127.0.0.1
port=3306Once the parameters are defined in the config file, you no longer need
to supply them in the dbConnect call, which then becomes much simpler:
Use the dbGetQuery function to submit your SQL to the database and
read the result sets. Doing so requires an open database connection:
You are not restricted to SELECT statements. Any SQL that
generates a result set is OK. It is common to use CALL statements, for
example, if your SQL is encapsulated in stored procedures and
those stored procedures contain embedded SELECT statements.
Using dbGetQuery is convenient because it packages the result set into
a data frame and returns the data frame. This is the perfect
representation of an SQL result set. The result set is a tabular data
structure of rows and columns, and so is a data frame. The result set’s
columns have names given by the SQL SELECT statement, and R uses them
for naming the columns of the data frame.
Call dbGetQuery repeatedly to perform multiple queries. When
you are done, close the database connection using dbDisconnect:
Here is a complete session that reads and prints three rows from a database of stock prices. The query selects the price of IBM stock for the last three days of 2008. It assumes that the username, password, dbname, and host are defined in the my.cnf file:
con <- dbConnect(RMySQL::MySQL())
sql <- paste(
"select * from DailyBar where Symbol = 'IBM'",
"and Day between '2008-12-29' and '2008-12-31'"
)
rows <- dbGetQuery(con, sql)
dbDisconnect(con)
print(rows)## Symbol Day Next OpenPx HighPx LowPx ClosePx AdjClosePx
## 1 IBM 2008-12-29 2008-12-30 81.72 81.72 79.68 81.25 81.25
## 2 IBM 2008-12-30 2008-12-31 81.83 83.64 81.52 83.55 83.55
## 3 IBM 2008-12-31 2009-01-02 83.50 85.00 83.50 84.16 84.16
## HistClosePx Volume OpenInt
## 1 81.25 6062600 NA
## 2 83.55 5774400 NA
## 3 84.16 6667700 NASee Also
See Recipe 3.10, “Installing Packages from CRAN”, and the documentation for RMySQL, which
contains more details about configuring and using the package.
See Recipe 4.17, “Accessing a Database with dbplyr”, for information about how to get data from an SQL without writing any SQL.
R can read from several other RDBMS systems, including Oracle, Sybase, PostgreSQL, and SQLite. For more information, see the R Data Import/Export guide, which is supplied with the base distribution (Recipe 1.7, “Viewing the Supplied Documentation”) and is also available on CRAN.
4.17 Accessing a Database with dbplyr
Problem
You want to access a database, but you’d rather not write SQL code in order to manipulate data and return results to R.
Solution
In addition to being a grammar of data manipulation, the tidyverse package dplyr can, in connection with the dbplyr package, turn dplyr commands into SQL for you.
Let’s set up an example database using RSQLite. Then we’ll connect to it and use dplyr and the dbplyr backend to extract data.
Set up the example table by loading the msleep example data into an in-memory SQLite database:
Now that we have a table in our database, we can create a reference to it from R:
The sleep_table object is a type of pointer or alias to the table on the database. However, dplyr will treat it like a regular tidyverse tibble or data frame, so you can operate on it using dplyr and other R commands. Let’s select all animals from the data who sleep less than three hours.
The dbplyr backend does not go fetch the data when we do the preceding commands. But it does build the query and get ready. To see the query built by dplyr, you can use show_query:
show_query(little_sleep)
#> <SQL>
#> SELECT *
#> FROM (SELECT `name`, `genus`, `order`, `sleep_total`
#> FROM `sleep`)
#> WHERE (`sleep_total` < 3.0)To bring the data back to your local machine, use collect:
Discussion
When you use dplyr to access SQL databases by writing only dplyr commands, you can be more productive by not having to switch from one language to another and back. The alternative is to have large chunks of SQL code stored as text strings in the middle of an R script, or have the SQL in separate files that are read in by R.
By allowing dplyr to transparently create the SQL in the background, you are freed from having to maintain separate SQL code to extract data.
The dbplyr package uses DBI to connect to your database, so you’ll need a DBI backend package for whichever database you want to access.
Some commonly used backend DBI packages are:
odbcUses the Open Database Connectivity (ODBC) protocol to connect to many different databases. This is typically the best choice when you are connecting to Microsoft SQL Server. ODBC is typically straightforward on Windows machines but may require some considerable effort to get working in Linux or Mac.
RPostgreSQLFor connecting to Postgres and Redshift.
RMySQLFor MySQL and MariaDB
RSQLiteFor connecting to SQLite databases on disk or in memory
bigrqueryFor connections to Google’s BigQuery.
Each DBI backend package discussed here is listed on CRAN and can be installed with the typical install.packages('*packagename*') command.
See Also
For more information about connecting the databases with R & RStudio, see https://db.rstudio.com/.
For more detail on SQL translation in dbplyr, see the sql-translation vignette at vignette("sql-translation") or http://dbplyr.tidyverse.org/articles/sql-translation.html.
4.18 Saving and Transporting Objects
Problem
You want to store one or more R objects in a file for later use, or you want to copy an R object from one machine to another.
Solution
Write the objects to a file using the save function:
Read them back using the load function, either on your computer or on any platform that supports R:
The save function writes binary data. To save in an ASCII format, use
dput or dump instead:
Discussion
We’ve found ourselves with a large, complicated data object that we want to load into other workspaces, or we may want to move R
objects between a Linux box and a Windows box. The load and save functions let us do all this: save will store the object in a file that is portable across machines, and load can read those files.
When you run load, it does not return your data per se; rather, it creates variables in your workspace, loads your data into those
variables, and then returns the names of the variables (in a vector). The first time you run load, you might be tempted to do this:
Let’s look at what myData is:
This might be puzzling, because myData will not contain your data at all. This can be perplexing and frustrating the first time.
The save function writes in a binary format to keep the file small.
Sometimes you want an ASCII format instead. When you submit a question
to a mailing list or to Stack Overflow, for example, including an ASCII dump of the data lets
others re-create your problem. In such cases use dput or dump, which
write an ASCII representation.
Be careful when you save and load objects created by a particular R
package. When you load the objects, R does not automatically load the
required packages, too, so it will not “understand” the object unless
you previously loaded the package yourself. For instance, suppose you
have an object called z created by the zoo package, and suppose we
save the object in a file called z.RData. The following sequence of
functions will create some confusion:
load("./data/z.RData") # Create and populate the z variable
plot(z) # Does not plot as expected: zoo pkg not loaded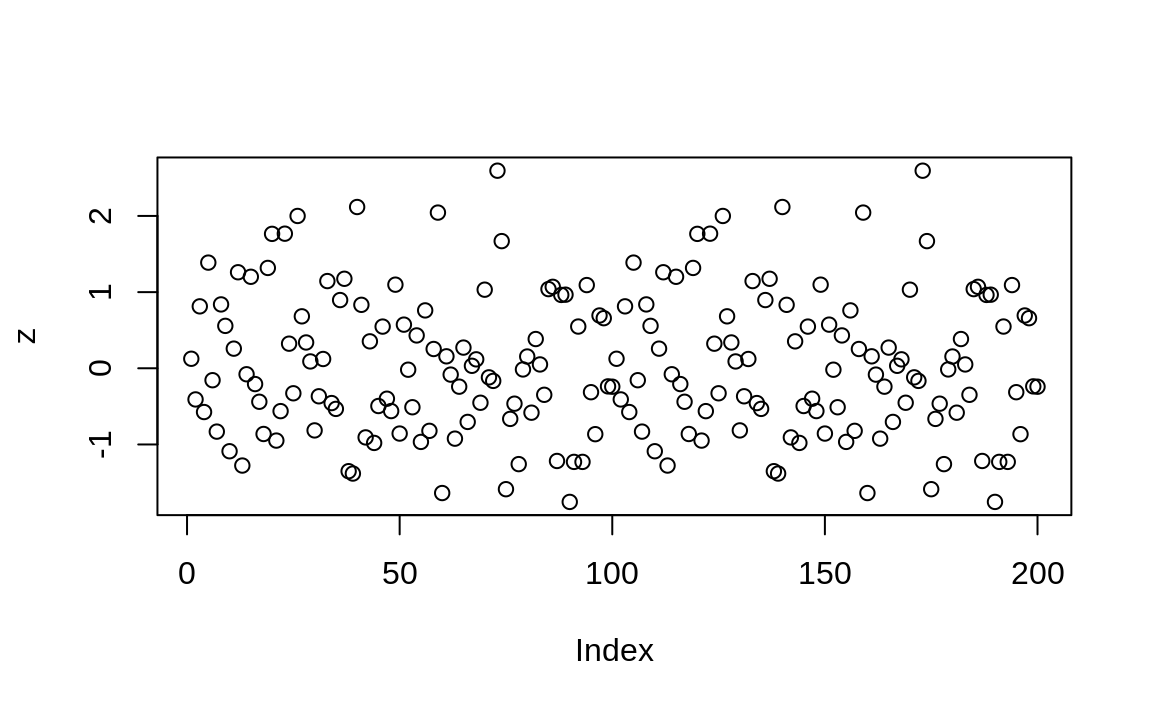
Figure 4.3: Plot without zoo loaded
The plot in Figure 4.3 shows the resulting plot, which is just points.
We should have loaded the zoo package before printing or plotting
any zoo objects, like this:
library(zoo) # Load the zoo package into memory
load("./data/z.RData") # Create and populate the z variable
plot(z) # Ahhh. Now plotting works correctly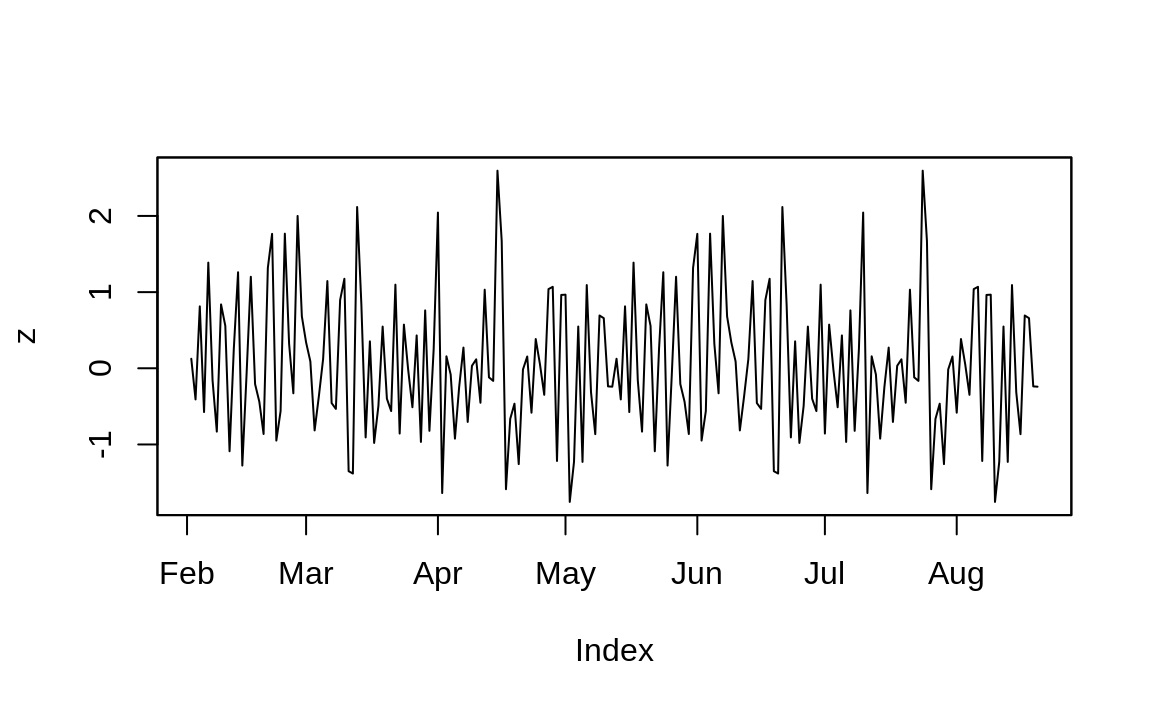
Figure 4.4: Plotting with zoo
And you can see the resulting plot in Figure 4.4.
See Also
If you are just saving and loading a single data frame or other R object, you should consider write_rds and read_rds. These functions don’t have “side effects” like load.